Previously undescribed lineage of Archaea illuminates microbial evolution
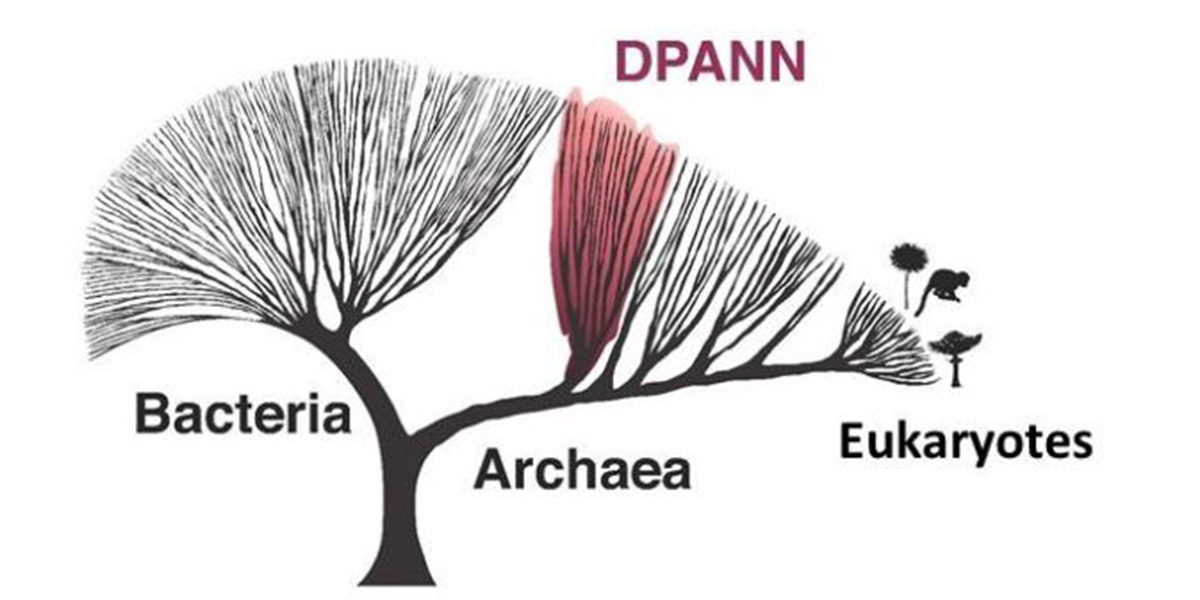
Archaea make up one of the main divisions of life, next to the Bacteria and the Eukaryotes, the latter of which comprise for example fungi, plants and animals. Archaea are a large group of microorganisms that live in all habitats on Earth ranging from soils and sediments to marine and freshwater environments as well as from human-made to host-associated habitats including the gut. In turn, Archaea are now thought to play a major role in biogeochemical nutrient cycles.
In a publication in Nature Communications last Friday, evolutionary microbiologists Nina Dombrowski and Anja Spang from the Royal Netherlands Institute of Sea Research (NIOZ) and Uppsala University (AS) describe a previously unknown archaeal lineage (phylum). The authors named them the Undinarchaeota, in reference to the female water spirit or nymph Undina. For the study, Dombrowski and Spang cooperated with partners from Uppsala and Bristol University, the University of Queensland and the Australian National University.
Diverse symbionts and parasites
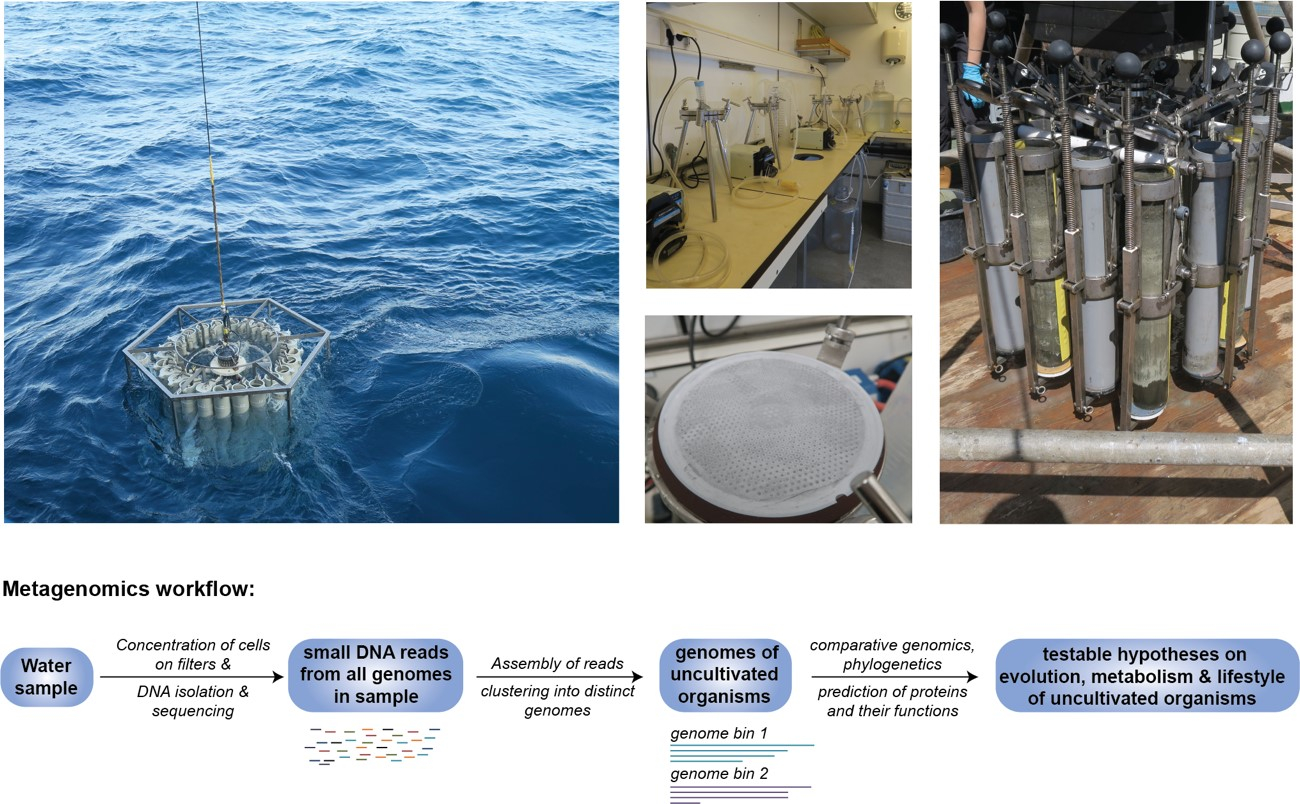
Because of their great resemblance to Bacteria, Archaea were only described as a separate lineage about 40 years ago and were not studied intensely until very recently, when it became possible to sequence DNA directly from environmental samples and to reconstruct genomes from uncultivated organisms. This field of genetic research, generally referred to as metagenomics (Figure 1), has not only revealed that microbial life including the Archaea is much more diverse than originally thought, but also provided data needed to shed light on the function of these microbes in their environments.
The newly described Undinarchaeota were discovered in genetic material from marine (Indian, Mediterranean and Atlantic ocean) and aquifer (Rifle aquiver, Colorado River) environments. The authors could show that they belong to a very diverse and until recently unknown group of so-called DPANN archaea. Members of the DPANN include organisms with very small genomes and limited metabolic capabilities, which suggests that these organisms depend on other microbes for growth and survival1,2,3. In fact, the few so far cultivated DPANN archaea are obligate symbionts or parasites that cannot live on their own4.
“In line with this, the Undinarchaeota seem to lack several anabolic pathways, indicating that they are, too, depend on various metabolites from so far unknown partner organisms”, says research leader Anja Spang. “However, Undinarchaeota seem to have certain metabolic pathways that lack in some of the most parasitic DPANN archaea and may be able to conserve energy by fermentation.”

Complex evolutionary history
While DPANN have only been discovered recently, it becomes increasingly clear that they are widespread and that representatives inhabit all thinkable environments on Earth. Yet, little is known about their evolutionary and ecological role. “In some way, some of the DPANN archaea resemble viruses, needing a host organism, likely other archaea or bacteria, for survival”, says Spang. “However, and in contrast to viruses, we currently know very little about the DPANN archaea and how they affect food webs and host evolution. It is also unclear whether DPANN are an ancient archaeal lineage that resembles early cellular life or have evolved later or in parallel with their hosts.”
With their study, the authors could shed more light on the complex evolution of Archaea. “Our work revealed that many DPANN archaea frequently exchange genes with their hosts, which makes it very challenging to reconstruct their evolutionary history”, says first author Nina Dombrowski. Tom Williams (Bristol University) adds: “However, we could show that DPANN have probably evolved in parallel with their hosts over a long evolutionary time scale, by identifying and studying those genes that were inherited from parent-to-offspring instead of having been transferred between host and symbiont”.
Role in marine biogeochemical cycles
Spang expects that certain DPANN including the Undinarchaeota, may be important for biogeochemical nutrient cycles within the oceans and sediments. “One reason that DPANN were discovered relatively recently, is that they were not retained on the filters originally used for concentrating cells from environmental samples due to their small cell sizes.” But since their discovery, DPANN turned out to be much more widespread than originally anticipated. Chris Rinke from the University of Queensland: “Prospective research on the Undinarchaeota and other DPANN archaea will be essential to obtain a better understanding of marine biogeochemical cycles and the role symbionts play in the transformation of organic matter.”
These questions drive some of the prospective projects of Anja Spang. In particular, in collaboration with their NIOZ colleagues Laura Villanueva, Pierre Offre and Julia Engelmann, the authors of the publication Anja Spang and Nina Dombrowski have just sequenced new DNA from water samples from the Black Sea, revealing that Undinarchaeota are present in almost all anoxic depth layers of this basin. Spang says: “These data are a gold mine for the future exploration of the ecology and evolution of these potentially symbiotic Archaea, allowing us to identify their interaction partners and to unravel further secrets about the biology of the Undinarchaeota.”
Figure 2. Depiction of the evolutionary placement of Undinarchaeota in an archaeal tree as a sister group of the symbiotic DPANN archaea (left). Illustration of cell of an Undinarchaeota representative, which might have pili (cell appendages) and may be able to take up DNA but is likely dependent on partner organisms for the acquisition of various metabolites (right).
Publication
Nina Dombrowski*, Tom A. Williams, Jiarui Sun, Benjamin J. Woodcroft, Jun-Hoe Lee, Bui Quang Minh, Christian Rinke & Anja Spang*
Undinarchaeota illuminate DPANN phylogeny and the impact of gene transfer on archaeal evolution
Nature Communications, 7 August 2020, https://doi.org/10.1038/s41467-020-17408-w
* Affiliated to Royal NIOZ and Utrecht University
References
1Castelle CJ, Wrighton KC, Thomas BC, et al. Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling. Curr Biol. 2015;25(6):690-701. doi:10.1016/j.cub.2015.01.014
2Rinke C, Schwientek P, Sczyrba A, et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature. 2013;499(7459):431-437. doi:10.1038/nature12352
3Dombrowski N, Lee JH, Williams TA, Offre P, Spang A. Genomic diversity, lifestyles and evolutionary origins of DPANN archaea. FEMS Microbiol Lett. 2019;366(2):fnz008. doi:10.1093/femsle/fnz008
4Huber H, Hohn MJ, Rachel R, Fuchs T, Wimmer VC, Stetter KO. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature. 2002;417(6884):63-67. doi:10.1038/417063a